| The main aim of this group is to analyse host-pathogen
interactions, at a molecular level. We aim to identify and functionally
characterise the bacterial effectors that carry out that interaction
within the host cell. Such effectors molecules are mostly secreted by
type III secretion systems, are abundant within different pathogens and
determine the nature of the infectious resistance and processes. Type III secretion systems are highly conserved. However, although some effectors can be common to different pathogens, the complete set of secreted effectors (secretome) is specific to each pathogen. The secretome determines which host cell processes are affected and how, and thus determine the effects of the infection. On this basis, the main interest of the group is the identification and functional characterisation of the effectors as well as their cellular targets, in order to understand which host processes are being affected by the pathogen and how. Currently, we are approaching this goal by two different basic strategies: A) Proteomic analysis of the secretome, and B) Identification of effectors on the basis of their secretion signals (functional genomics). This research project is funded by the National Plan for Biotechnology (Ministry of Education and Science, Spain). To carry out this project, the experience, methods and tools acquired and developed by the principal investigator during her previous work on identification and functional characterisation of TTSS in Salmonella is being transferred to the analysis of the interaction between Pseudomonas syringae and the host. P. syringae includes many pathovars that affect a wide variety of commercially important crops and has been selected as a model for being a well establish and representative model, and because it expresses a TTSS that is essential for its pathogenesis. |
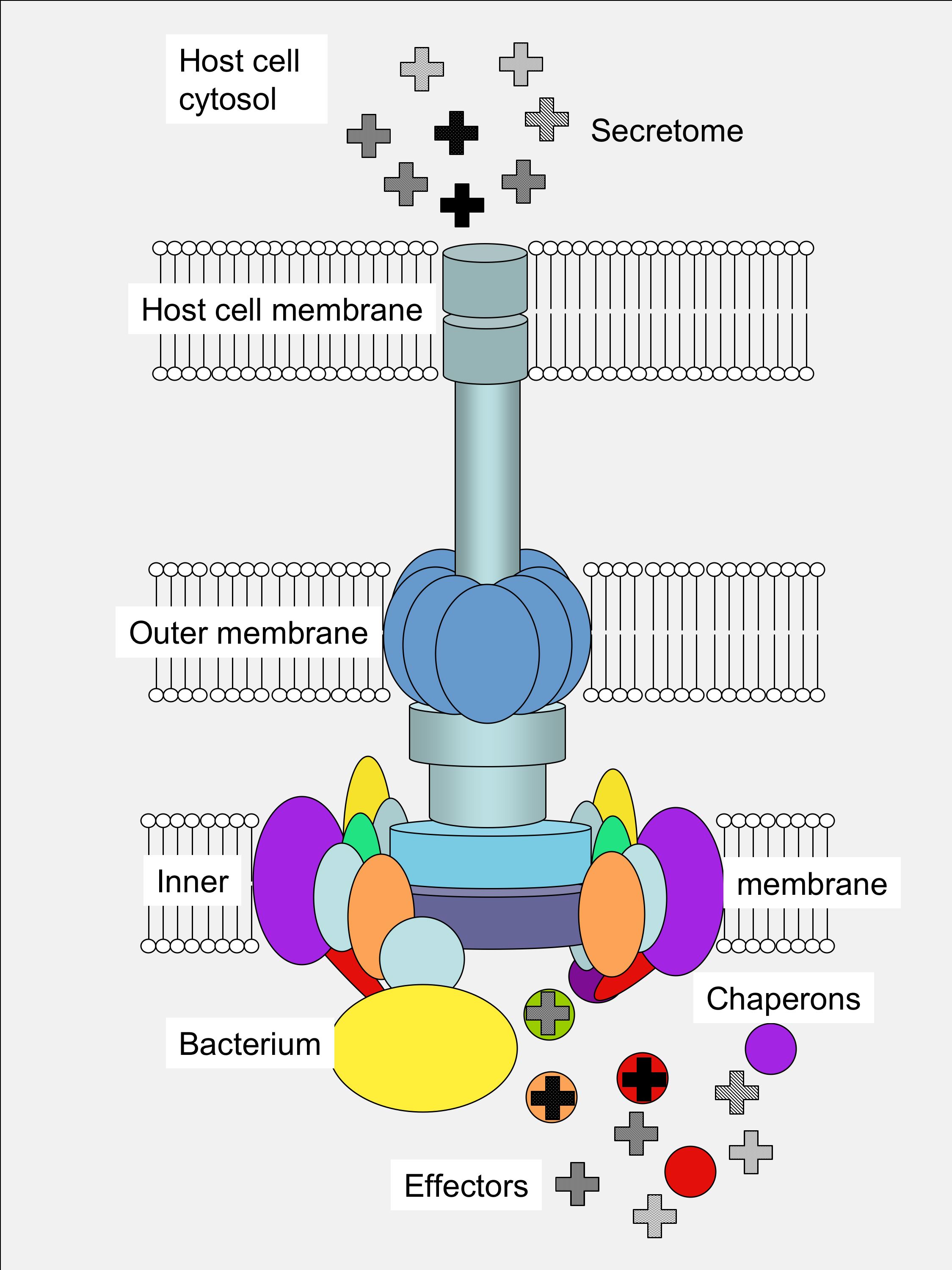 Type III secretion system structure
Bean plants
Infected with
Control
P. syringae pv. phaseolicola |